The proposed MASTER project will take place within the Computational Lung working group (MYRIAD team) which focuses on the acute respiratory distress syndrome (ARDS) to help intensive care clinicians by using computed tomography (CT) acquired at different respiratory conditions (e.g., end-inhale and end-exhale). Changes between CT scans can be used to assess such phenomena as alveolar recruitment or cyclic hyperinflation. The question that we raise is whether (or not) a novel self-supervised training strategy based on both end-inhale and end-exhale images can improve CT-lung image analysis (segmentation or registration) compared to using only one respiratory condition, i.e., improving the latent space embedding by taking advantage of having a pair of images.
- The objective of this master project is to enhance the encoding of thoracic CT-scans by developping a novel self-supervised deep representation based on double respiratory CT imaging of the patient.
- The candidate will train deep models on several hundreds of thoracic CT-scans gathered from several databases among them the COVID database acquired in CREATIS during last year.
- The developed deep models and self-supervised strategies will be specifically designed to handle the challenging specificity of having large-size 3-D medical data and data of the same patient acquired at several respiratory phases.
- Finally, the comparison between several self-supervised approaches will yield the best pre-training strategy to improve the performance of several deep models dedicated to CT-lungs analysis (segmentation, registration, ...).
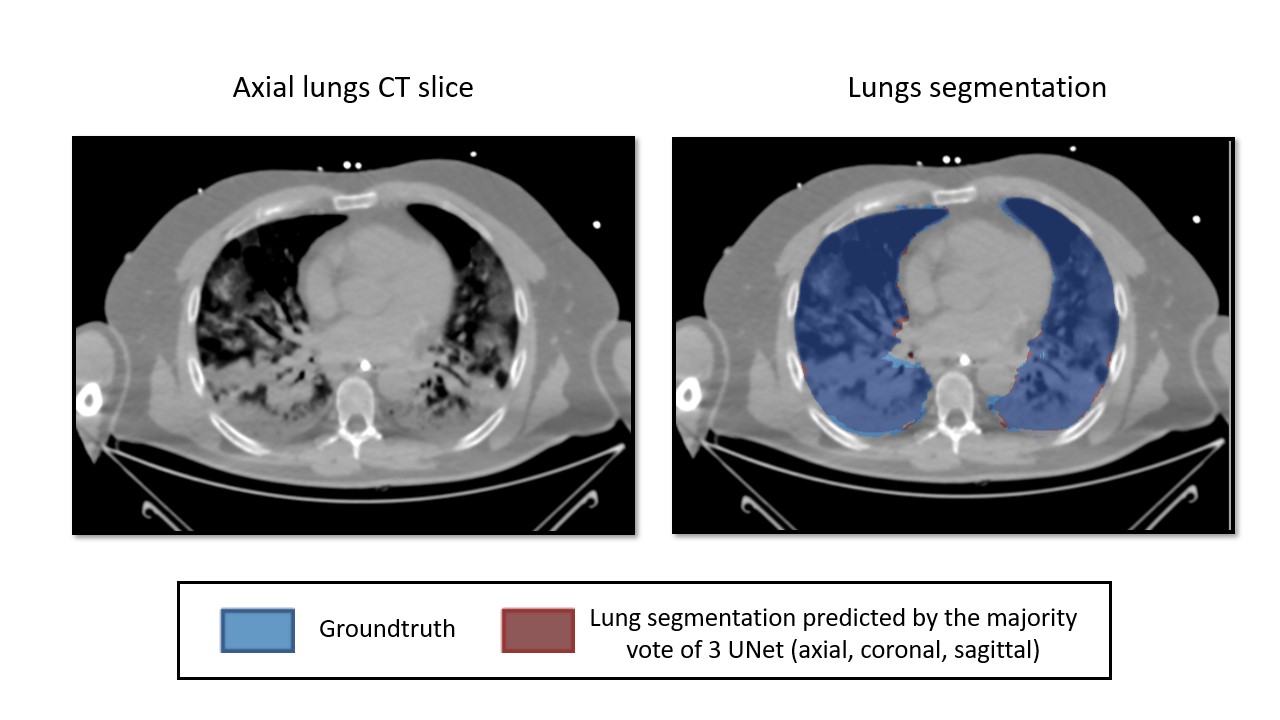
For instance, you can find hereafter an example illustrating the issues raised by the very high opacities (ARDS) in a lung segmentation task :