Abstract
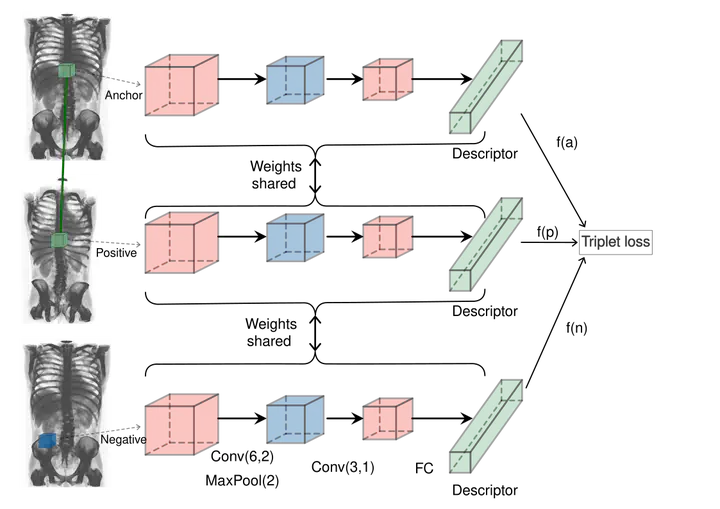
Purpose: We propose to learn a 3D keypoint descriptor which we use to match keypoints extracted from full-body CT scans. Our methods are inspired by 2D keypoint descriptor learning, which was shown to outperform hand-crafted descriptors. Adapting these to 3D images is challenging because of the lack of labelled training data and high memory requirements. Method : We generate semi-synthetic training data. For that, we first estimate the distribution of local affine inter-subject transformations using labelled anatomical landmarks on a small subset of the database. We then sample a large number of transformations and warp unlabelled CT scans, for which we can subsequently establish reliable keypoint correspondences using guided matching. These correspondences serve as training data for our descriptor, which we represent by a CNN and train using the triplet loss with online triplet mining. Results : We carry out experiments on a synthetic data reliability benchmark and a registration task involving 20 CT volumes with anatomical landmarks used for evaluation purposes. Our learned descriptor outperforms the 3D-SURF descriptor on both benchmarks while having a similar runtime. Conclusion: We propose a new method to generate semi-synthetic data and a new learned 3D keypoint descriptor. Experiments show improvement compared to a hand-crafted descriptor. This is promising as literature has shown that jointly learning a detector and a descriptor gives further performance boost.