inTag : Cardiac MRI tagging analysis toolbox - Introduction
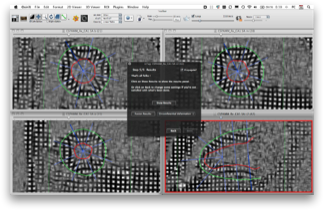
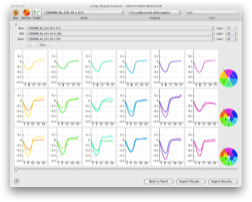
inTag is a software to calculate, display and analyze myocardial strains and intra-myocardial mechanics from cardiac MR images with a tagging pattern.
inTag brings processing and analysis of cardiac tagged MR sequences to most in the clinical environment. One of the main reasons for MR tagging not to be more frequently used in preclinical of clinical research, is the lack of widely available processing tools.
inTag brings processing and analysis of cardiac tagged MR sequences to most in the clinical environment. One of the main reasons for MR tagging not to be more frequently used in preclinical of clinical research, is the lack of widely available processing tools.
inTag offers a fast and integrated process that bring quantitative strain analysis to most scientists, or physicians in a matter of minutes.
Motion estimation in InTag is based on the Sine Wave Modeling approach (SinMod) (1). With this method, image intensity in the environment of each pixel is modeled by a moving sine wave front. Displacement is estimated at sub-pixel accuracy (displacement errors <0.02 pixels). In a comparison with the HARP method (1), SinMod compares favorably and appears less sensitive to artifacts, especially later in the cardiac cycle.
(1) Mapping displacement and deformation of the heart with local sine wave modeling. T.Arts, F.W.Prinzen, T.Delhaas, J.Milles, A.Rossi, P.Clarysse. IEEE Trans Med Imag 2010 May;29(5):1114-23
(1) Mapping displacement and deformation of the heart with local sine wave modeling. T.Arts, F.W.Prinzen, T.Delhaas, J.Milles, A.Rossi, P.Clarysse. IEEE Trans Med Imag 2010 May;29(5):1114-23
After a preliminary prototyping step in Matlab ®, InTag is now available as a plugin in OsiriX, the advanced open-source image processing software. It benefits of all its unique capabilities for navigation, visualisation of multimodality and multidimensional images, and data management.
The Matlab to OsiriX environment transfer was performed by Axinoe and this work was funded by a French Government grant and Hôpitaux de Lyon (HCL).
The Matlab to OsiriX environment transfer was performed by Axinoe and this work was funded by a French Government grant and Hôpitaux de Lyon (HCL).